Cytoscape software6/23/2023 Modularization method family offering modular hierarchies and adjustable overlaps. The Java Universal Network/Graph Framework-is a software library that provides a common and extendible language for the modeling, analysis, and visualization of data that can be represented as a graph or network.Ĭomputes group centrality scores and identifies the most central group of players in a network. Software for Social Network Analysis & Organizational Network Analysis.Ī tool for virtual experimental network topological analysis. GTNA is a Java-based framework that allows for the graph-theoretic analysis of arbitrary network topologies. The core data structures and algorithms are implemented in C++. Graph-tool is an efficient Python module for manipulation and statistical analysis of graphs (a.k.a. You can generate, import, export, measure, layout and visualize them. GraphStream is a Java library for the modeling and analysis of dynamic graphs. Node betweenness centrality (parallelized)ĭisk-based large-scale graph computation. Is a MATLAB toolbox for inference and analysis of biological and cellular networks. It helps the user to collect and analyse all the egocentric network data (all social network data of a website on the Internet), and provide general global network measures and data matrixes that can be used for further analysis by other software. Local Average Connectivity-based method (LAC)ĮgoNet (Egocentric Network Study Software) for the collection and analysis of egocentric social network data. Providing calculation, evaluation and visualization analysis for several centralities of weighted and unweighted network. It is a tool for centrality comparison using dimensional reduction algorithmsĬytoHubba is a Java plugin for Cytoscape, a facilitated platform for the analysis and visualization of molecular interaction networks based on web application, Hubba.ĭensity of Maximum Neighborhood Component (DMNC)ĭouble Screening Scheme (DSS) of MNC || DMNC It is an R package available on CRAN repository specialized for network centrality analysis. Use pathway topology information to assign weight to pathway nodes. Shannon-entropy based diversity coefficientĬentiLib is a Java-library for the computation and investigation of weighted and unweighted centralities in biological networks.įind the most important nodes in a network, calculating centrality parameters for each node.Ĭentrality-based pathway enrichment. These measures are increasingly used to characterize structural and functional brain connectivity datasets. 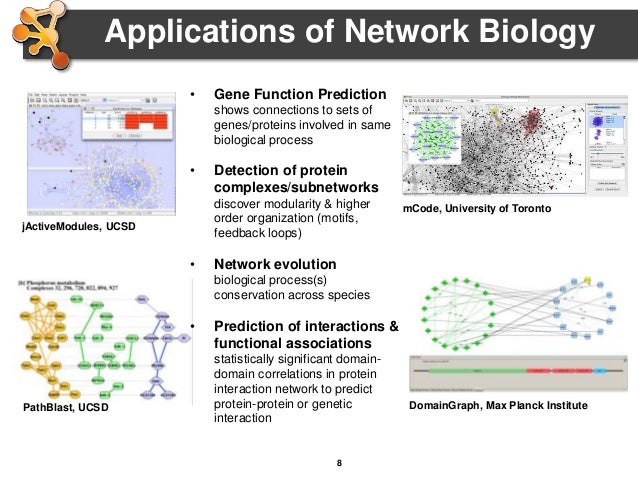
The BCT contains a large selection of complex network measures in Matlab. AllegroGraph supports SPARQL, RDFS++, and Prolog reasoning from numerous client applications. 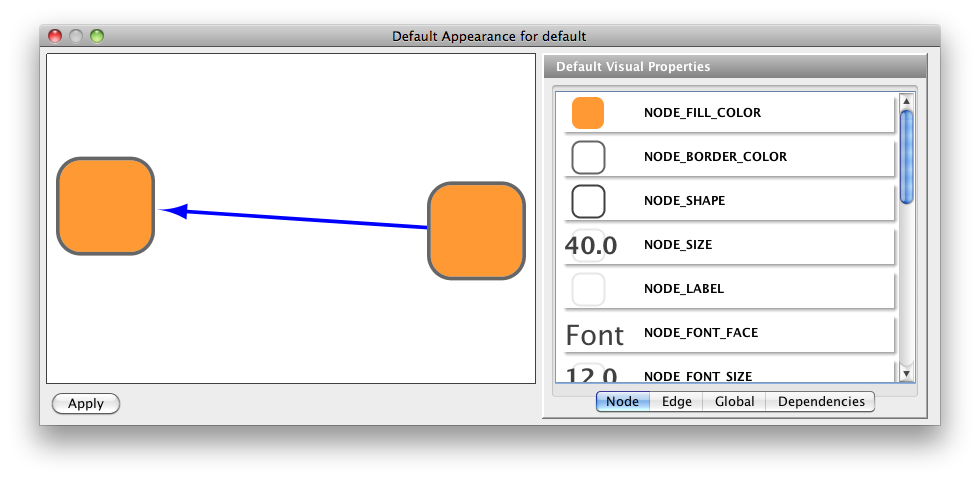
AllegroGraph uses efficient memory utilization in combination with disk-based storage, enabling it to scale to billions of quads while maintaining superior performance. AllegroGraph® is a modern, high-performance, persistent graph database.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
